Infectious Diseases
Should Not Kill Anyone
Universal Infectious Diseases Test (UID)
A Revolutionary Technology to Aid the Clinician
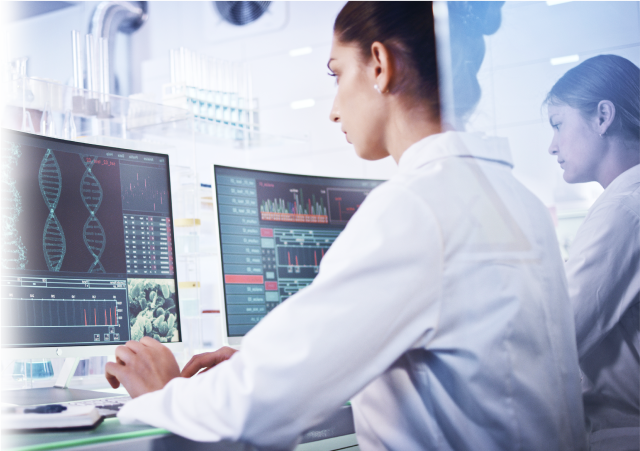

Sepsis And Its Global Impact
Sepsis is the body's extreme response to an infection and is potentially life-threatening. This occurs when a pre-existing infection
triggers a chain reaction throughout your body, often leading to shock, disability, multi organ failure, or even death.
SEPSIS IS A LEADING CAUSE OF DEATH
- Sepsis 11Mn
- TB 1.5Mn
- Cancer 10Mn
- Stroke 6Mn
>2 BILLION PEOPLE WITH THE FOLLOWING PRE-EXISTING
CONDITIONS ARE AT A HIGHER RISK OF SEPSIS
The Universal Infectious Diseases Test
UID Test is an NGS based culture-free test to identify causative pathogen in a quick turn around time
1200+ PATHOGENS
Comprehensive single screening test covering bacteria and fungi
ARG PROFILING
Identifies drug resistance based on Antibiotic Resistance Gene (ARG) profile
RESULT REPORTED IN < 12 HOURS
Hands on time < 4 hours
The Current System Needs An Upgrade To Save Lives
| Tests for ID | Turn Around Time | Pathogen Coverage | Antibiotic Coverage | Additional Information |
|---|---|---|---|---|
12 hours | >1200 pathogens | ARGs | Species and Genus identification | |
2-10 Days | Culturable bacteria and fungi | Selected Panel | ||
12-24 Hours | Up to 43 pathogens | 1-5 Antibiotics | Upgradation of technology on existing set up is not possible or is very difficult |
Our Partners
100+
CITIES
100+
HOSPITALS
500+
DOCTORS
How may we help you?
In the news
24 April 2022 | The Times of India
COVID-19 led to a spike in Tuberculosis cases in India: Experts
24 March 2022 | Medgate today Magazine
Early Detection and Accurate Diagnosis crucial to make India TB Free by 2025 - HaystackAnalytics Insights
7 February 2022 | The Economic Times
HaystackAnalytics develops first Universal Infectious Disease (ID) genomic test

22 January 2022 | Financial Express
Public Health: Tech a formidable ally against TB





















